from matplotlib.pylab import f
import numpy as np
# Let's compare MU and Bregman mirror descent on a simple Poisson regression problem
# Hyperparameters
m, n = 100, 20 # number of samples and features
alpha = 100 # Poisson signal level
hgt = np.maximum(np.random.randn(n),0)
W = np.random.rand(m,n)
#W = 0.5*W*np.ones(n) + 0.5*W
y = np.random.poisson(alpha * W @ hgt)
# Petit check norme spectrale
nW = np.linalg.norm(W, 2)
print(f'Spectral norm of W: {nW}')
gamma = 1e-1
tau = 0.99/nW/gamma
print("default value of tau:", tau)
print("default value of gamma:", gamma)
# Bach Heuristic
#alpha = np.sum(y)/np.sum(W)
#tau = np.sqrt(m/n)/alpha/nW
#gamma = np.sqrt(n/m)*alpha/nW
#print(f"Using tau={tau} and gamma={gamma} from Yanez-Bach heuristic")
def loss(h): # KL divergence
return np.sum(W@h-y) + np.sum(y * np.log((y+1e-10)/(W @ h)))
def MU(y, W, h, itermax=1000, epsilon=1e-10):
Wtsum = W.T @ np.ones_like(y) # suboptimal but ok
hout = np.copy(h)
loss_val = [loss(hout)]
for i in range(itermax):
hout = np.maximum(epsilon, hout * (W.T @ (y / (W @ hout ))) / (Wtsum))
loss_val.append(loss(hout))
return hout, loss_val
def BurgMU(y, W, h, itermax=1000, epsilon=1e-10): # NoLIPS by Bauschke et al
Wtsum = W.T @ np.ones_like(y) # suboptimal but ok
lamb = 1/2/np.sum(y)
hout = np.copy(h)
loss_val = [loss(hout)]
for i in range(itermax):
hout = np.maximum(epsilon, hout / ( 1 + lamb* hout * ( Wtsum - W.T @ (y / (W @ hout )))))
loss_val.append(loss(hout))
return hout, loss_val
def prox_kl(y, x, tau):
return 0.5 * (x - tau + np.sqrt((x - tau)**2 + 4 * tau * y))
def prox_kl_dual(y, x, tau):
return 0.5 * (x - np.sqrt(x**2 + 4 * tau * y))
# Condat-Vu setting
# min eta_+(h) + KL(y, Wh)
# L = W
# f = eta_+
# g = KL(y, .)
# use prox_tau*g* = Id - tau*prox_kl(y, ./tau, 1/tau)
def CondatVu(y, W, h, itermax=1000, epsilon=1e-10, tau=None, gamma=1e-3):
hout = np.copy(h)
if tau is None:
tau = 0.99/gamma/nW
print(f"Using tau={tau}")
u = np.zeros_like(W@h) # dual variable
loss_val = [loss(hout)]
for i in range(itermax):
uold = np.copy(u)
utilde = u + tau*W@hout
u = utilde - tau*prox_kl(y, utilde/tau, 1/tau)
hout = np.maximum(epsilon, hout - gamma*W.T@(2*u - uold))
loss_val.append(loss(hout))
return hout, loss_val
def CondatVuAlt(y, W, h, itermax=1000, epsilon=1e-10, tau=None, gamma=1e-3):
hout = np.copy(h)
if tau is None:
tau = 0.99/gamma/nW
print(f"Using tau={tau}")
u = np.zeros_like(W@h) # dual variable
loss_val = [loss(hout)]
for i in range(itermax):
utilde = u + tau*W@hout
u = utilde - tau*prox_kl(y, utilde/tau, 1/tau)
hout = np.maximum(epsilon, hout - gamma*W.T@u)
loss_val.append(loss(hout))
return hout, loss_val
def CondatVuAltLoop(y, W, h, itermax=1000, epsilon=1e-10, tau=None, gamma=1e-3):
hout = np.copy(h)
if tau is None:
tau = 0.99/gamma/nW
print(f"Using tau={tau}")
u = np.zeros_like(W@h) # dual variable
loss_val = [loss(hout)]
for i in range(itermax):
Wh = W@hout
for _ in range(5): # inner loop
utilde = u + tau*Wh
u = utilde - tau*prox_kl(y, utilde/tau, 1/tau)
Wtu = W.T@u
for _ in range(5):
hout = np.maximum(epsilon, hout - gamma*Wtu)
loss_val.append(loss(hout))
return hout, loss_val
def ChambollePockBach(y, W, hinit=None, itermax=1000, epsilon=1e-10, gamma=None, tau=None):
hout = np.copy(hinit)
if tau is None and gamma is None:
# Bach Heuristic
alpha = np.sum(y)/np.sum(W)
tau = np.sqrt(m/n)/alpha/nW
gamma = np.sqrt(n/m)*alpha/nW
print(f"Using tau={tau} and gamma={gamma}")
loss_val = [loss(hout)]
u = np.zeros_like(W@hout) # W@hout
htilde = np.copy(hout)
hold = np.copy(hout)
for i in range(itermax):
utilde = u + tau*W@htilde # TODO why not - like in paper??
u = (utilde - np.sqrt(utilde**2 + 4*tau*y))/2
hout = np.maximum(epsilon, hout - gamma*W.T@(u + 1))
htilde = 2*hout - hold
hold = np.copy(hout)
loss_val.append(loss(hout))
return hout, loss_val
hmu, loss_mu = MU(y, W, alpha*np.ones(n), itermax=5000)
hburg, loss_burg = BurgMU(y, W, alpha*np.ones(n), itermax=5000)
hpd, loss_pd = CondatVu(y, W, alpha*np.ones(n), itermax=5000, tau=tau, gamma=gamma)
hpda, loss_pda = CondatVuAlt(y, W, alpha*np.ones(n), itermax=5000, tau=tau, gamma=gamma)
hpdal, loss_pdal = CondatVuAltLoop(y, W, alpha*np.ones(n), itermax=5000, tau=tau, gamma=gamma)
#hcp, loss_cp = ChambollePockBach(y, W, alpha*np.ones(n), itermax=5000, tau=None, gamma=None) # very fast !
hcp, loss_cp = ChambollePockBach(y, W, alpha*np.ones(n), itermax=5000, tau=tau, gamma=gamma) # very fast !
import matplotlib.pyplot as plt
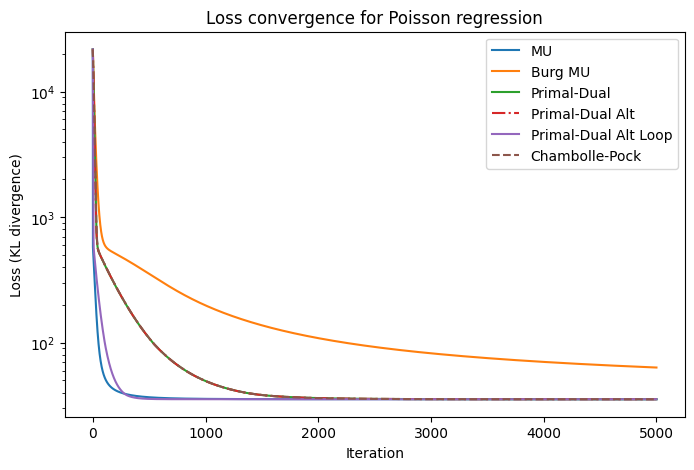
plt.figure(figsize=(8,5))
plt.semilogy(loss_mu, label='MU')
plt.semilogy(loss_burg, label='Burg MU')
plt.semilogy(loss_pd, label='Primal-Dual')
plt.semilogy(loss_pda, label='Primal-Dual Alt', linestyle='-.')
plt.semilogy(loss_pdal, label='Primal-Dual Alt Loop')
plt.semilogy(loss_cp, label='Chambolle-Pock', linestyle='--')
plt.title('Loss convergence for Poisson regression')
plt.xlabel('Iteration')
plt.ylabel('Loss (KL divergence)')
plt.legend()
plt.show()
# Printing the final estimation error
print(f"Final loss MU: {loss_mu[-1]}, Burg MU: {loss_burg[-1]}, Primal-Dual: {loss_pd[-1]}, Primal-Dual Alt: {loss_pda[-1]}, Primal-Dual Alt Loop: {loss_pdal[-1]}, Chambolle-Pock: {loss_cp[-1]}")
print(f"Estimation error MU: {np.linalg.norm(hmu - alpha*hgt)/np.linalg.norm(alpha*hgt)}, Burg MU: {np.linalg.norm(hburg - alpha*hgt)/np.linalg.norm(alpha*hgt)}, Primal-Dual: {np.linalg.norm(hpd - alpha*hgt)/np.linalg.norm(alpha*hgt)}, Primal-Dual Alt: {np.linalg.norm(hpda - alpha*hgt)/np.linalg.norm(alpha*hgt)}, Chambolle-Pock: {np.linalg.norm(hcp - alpha*hgt)/np.linalg.norm(alpha*hgt)}")